How DICER Cleaves microRNAs with Precision
Breakthrough in RNA Silencing Research
A research team from The Hong Kong University of Science and Technology (HKUST) has made a significant breakthrough in understanding the molecular machinery behind RNA silencing. Their work has shed light on how the human enzyme DICER achieves highly precise processing of microRNAs (miRNAs), offering new insights into gene regulation and potential implications for cancer, immune disorders, and genetic diseases.
The study, led by Prof. Tuan Anh Nguyen, Associate Professor in the Division of Life Science at HKUST, was conducted jointly with his Ph.D. students Minh Khoa Ngo and Cong Truc Le. It has been published in Nature under the title “DICER cleavage fidelity is governed by 5′-end binding pockets.”
Understanding the Role of RNA
In human life, the message is encoded in genomic DNA through the transcription of messenger RNAs, which carry out the genetic instructions. RNA molecules, typically single-stranded and composed of ribonucleotides (A, U, G, and C), play essential roles in cellular processes such as protein synthesis, gene regulation, and serving as genetic material in certain viruses.
Within this complex RNA-based regulatory landscape, the enzyme DICER acts as a highly precise “molecular scissors.” It cleaves double-stranded RNA precursors into short regulatory RNAs that are then incorporated into the RNA-induced silencing complex (RISC). This allows the cell to identify and suppress incorrect or unnecessary genetic messages—similar to marking and deleting errors in a text.
Deciphering DICER’s Precision
For years, researchers have sought to understand how DICER achieves its remarkable cleavage accuracy. Using advanced biochemical and structural biology techniques, including high-resolution cryoelectron microscopy (cryo-EM), the HKUST research team visualized DICER–RNA interactions at an atomic level.
Minh Khoa Ngo, the study’s first author, explained, “CryoEM allowed us to observe how RNA substrates engage with DICER at an atomic detail. These structural snapshots vividly reveal the dynamic adjustments DICER makes when processing different RNA sequences, fundamentally reshaping our understanding of how this enzyme functions.”
Key Findings and Mechanisms
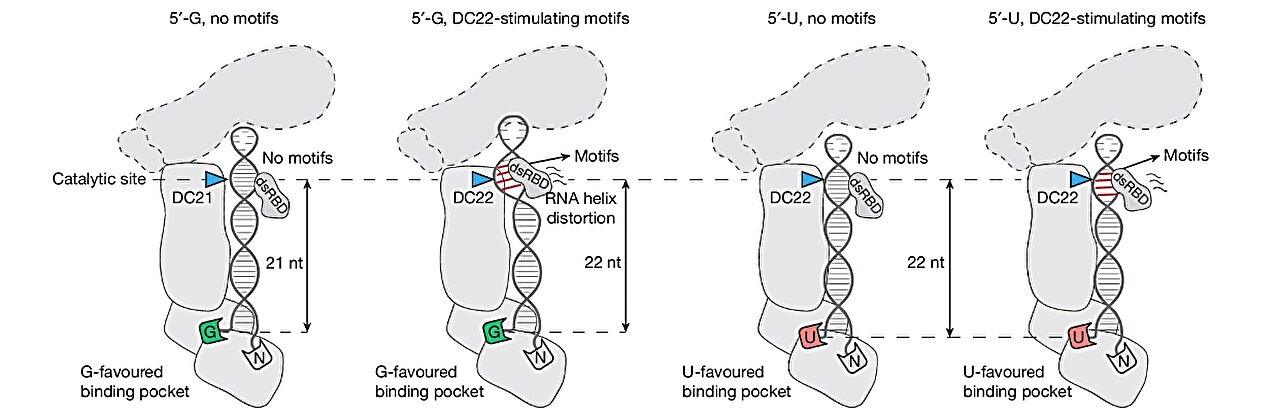
The team discovered that before cleavage occurs, DICER undergoes conformational adjustments that guide RNA substrates into the correct register. The enzyme then uses specific structural elements—particularly amino acids within 5′-end binding pockets—to align the RNA precisely before adopting the “cleavage ready” conformation.
Prof. Nguyen, the corresponding author, elaborated, “It is as if the scissors can ‘read’ exactly where the RNA should be cut at single nucleotide resolution, ensuring the integrity of the entire message. Our study uncovers not only the previously known U-favored 5′-end binding pocket, but also a newly identified G-favored 5′-end binding pocket.”
“Together, these form a dual-pocket mechanism that determines cleavage positioning, providing an entirely new framework for understanding how DICER accommodates and processes diverse RNA substrates.”
Implications Beyond Basic Biology
The importance of this discovery extends beyond basic biology. By revealing how DICER integrates 5′-end identity, RNA motifs, and domain motions to maintain cleavage fidelity, the findings lay a mechanistic foundation for improving RNA-based therapeutics, optimizing gene silencing technologies, and uncovering the molecular origins of DICER-related genetic diseases.
Future Directions
This groundbreaking research opens up new possibilities for advancing gene regulation studies and developing targeted therapies. As scientists continue to explore the intricacies of DICER and its role in RNA processing, the potential applications in medicine and biotechnology could be vast and transformative.
The study, titled “DICER cleavage fidelity is governed by 5′-end binding pockets,” was published in Nature with the DOI: 10.1038/s41586-026-10211-5.
- Generasi Muda Tangguh, Fondasi Indonesia Menggapai Dunia - April 7, 2026
- How DICER Cleaves microRNAs with Precision - April 7, 2026
- Dr. Daniel, Apakah Konsultasi Diperlukan untuk Mengetahui Posisi Rahim Normal atau Terbalik? - April 7, 2026



Leave a Reply